There will be a document (perhaps several) based on the meeting, but until then here are a few quick thoughts. Note that the comments below are my own and you shouldn't read into this anything about what directions the GBIC document(s) will actually take.
Microbiology rocks
Highlight of the first day was Robert J. Robbin's talk which urged the audience to consider that life was mostly microbial, that the the things most people in the room cared about were actually merely a few twigs on the tree of life, that the tree of life didn't actually exist anyway, and many of the concepts that made sense for multicellular organisms simply didn't apply in the microbial world. Basically it was a homage to Carl Woese (see also Pace et al. 2012 doi:10.1073/pnas.1109716109) and a wake up call to biodiversity informaticians to stop viewing the world through multicellular eyes. (You can find all the keynotes from the first day here).
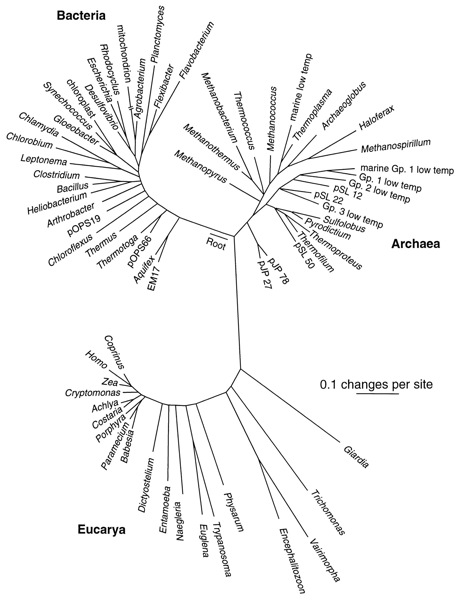
From Pace, N. R. (1997). A Molecular View of Microbial Diversity and the Biosphere. Science, 276(5313), 734–740. doi:10.1126/science.276.5313.734
Sequences rule
The future of a lot of biodiversity science belongs to sequences, from simple DNA barcoding as a tool for species discovery and identification, metabarcoding as a tool for community analysis, to comparisons of metabolic pathways and beyond. The challenge for classical biodiversity informatics is how to engage with this, and to what extent we should try and map between, say sequences and classical taxa, or whether it might make more sense (gasp) to abandon the taxonomic legacy and move on. Perhaps are more nuanced response is that the point of connection between sequences and classical biodiversity data is unlikely to be at the level of taxonomic names (which are mostly tags for collections of things that look similar) but at the level of specimens and observations.
Ontologies considered harmful
This is my own particular hobby horse. Often the call would come "we need an ontology", to which I respond read Ontology is Overrated: Categories, Links, and Tags. I have several problems with ontologies. The first is that they are too easy to make and distract from the real problem. From my perspective a big challenge is linking data together, that is going from

to
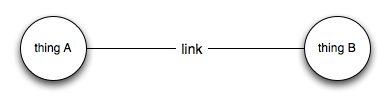
Let's leave aside what "A" and "B" are (I suspect it matters less than people think), once we have the link then we can can start to do stuff. From my perspective, what ontologies give us is basically this:
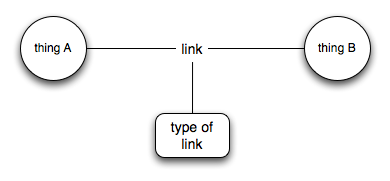
So now we know the "type" of the link (e.g., "is a part of", "cites approvingly", etc.). I'm not arguing that this isn't useful to have, but if you don't have the network of links then typing the links becomes an idle exercise.
To give an example, the web itself can be modelled as simply nodes connected by links, ignoring the nature of the links between the web pages. The importance of those links can be inferred later from properties of the network. To a first approximation this is how Google works, it doesn't ask what the links "mean" it simply investigates the connections to determine how important each web page is. In the same way, we build citation networks without bothering to ask the nature of the citation (yes I know there are ontologies for citations, but anyone willing to bet how widely they'll be adopted?).
My second complaint is that building ontologies is easy, "easy" in the sense that get a bunch of people together, they squabble for a long time about terminology, and out comes an ontology. Maybe, if you're lucky, someone will adopt it. The cost of making ontologies, and indeed of adopting them is relatively low (although it might not seem like it at the time). The cost of linking data is, I'd argue, higher, because it requires that you trust someone else's identifiers to the extent that you use them for things you care about deeply. Consider the citation network that is emerging from the widespread adoption of DOIs by the publishing industry. Once people trust that the endpoints of the links will survive, then the network starts to grow. But without that trust, that leap of faith, there's no network (unless you have enough resources to build the whole thing internally yourself, which is what happened with the closed citation network owned by Thomson Reuters). It's much easier to silo the data using unique identifiers than it is to link to other data (it's a variant of the "not invented here" syndrome).
Lastly, ontologies can have short lives. They reflect a certain world view that can become out of date, or supplanted if the relationships between things that the ontology cares about can be computed using other data. For example, biological taxonomy is a huge ontology that is rapidly being supplanted by phylogenetic trees computed from sequence (and other) data (compare the classification used by flagship biodiversity projects like GBIF and EOL with the Pace tree of life shown above). Who needs an ontology when you can infer the actual relationships? Likewise, once you have GPS the value of a geographic ontology (say of place names) starts to decline. I can compute if I'm on a mountain simply by knowing where I am.
I'm not saying ontologies are always bad (they're not), nor that they can't be cool (they can be), I'm just suggesting that they aren't the first thing you need. And they certainly aren't a prerequisite for linking stuff together.
Google flu trends
Perhaps the most interesting idea that emerged was the notion of intelligently detecting changes in biodiversity (which is the kind of thing a lot of people want to know) in the way analogous to Google.org's Flu Trends uses flu-related search terms to predict flu outbreaks:
Ginsberg, J., Mohebbi, M. H., Patel, R. S., Brammer, L., Smolinski, M. S., & Brilliant, L. (2008). Detecting influenza epidemics using search engine query data. Nature, 457(7232), 1012–1014. doi:10.1038/nature07634
Could we do something like this for biodiversity data? For various reasons this suggestion become known at GBIC2012 as the "Heidorn paradigm".
Thinking globally
One challenge for a meeting like GBIC 2012 is scope. There's so much cool stuff to think about. From my perspective, a useful filter is to ask "what will happen anyway?" In other words, there is a lot of stuff (for example the growth of metabarcoding) that will happen regardless of anything the biodiversity informatics community does. People will make taxon-specific ontologies for organismal traits, digitise collections, assess biodiversity, etc. without necessarily requiring an entity like GBIF. The key question is "what won't happen at a global scale unless GBIF (or some other entity) gets involved?"
A Vast Machine
 Lastly, in one session Tom Moritz mentioned a book that he felt we could learn from (A Vast Machine: Computer Models, Climate Data, and the Politics of Global Warming). The book recounts the history of climatology and its slow transition to a truly global science. I've started to read it, and it's fascinating to see the interplay between early visions of the future, and the technology (typically driven by military or large-scale commercial interests) that made possible the realisation of those visions. This is one reason why predicting the future is such a futile activity, the things that have the biggest effect come from unexpected sources, and effect things in ways it's hard to anticipate. On a final note, it took about a minute from the time from the time Tom mentioned the book to the time I had a copy from Amazon in the Kindle app on my iPad. Oh that accessing biodiversity data were that simple.
Lastly, in one session Tom Moritz mentioned a book that he felt we could learn from (A Vast Machine: Computer Models, Climate Data, and the Politics of Global Warming). The book recounts the history of climatology and its slow transition to a truly global science. I've started to read it, and it's fascinating to see the interplay between early visions of the future, and the technology (typically driven by military or large-scale commercial interests) that made possible the realisation of those visions. This is one reason why predicting the future is such a futile activity, the things that have the biggest effect come from unexpected sources, and effect things in ways it's hard to anticipate. On a final note, it took about a minute from the time from the time Tom mentioned the book to the time I had a copy from Amazon in the Kindle app on my iPad. Oh that accessing biodiversity data were that simple.